P. Lynne Howell
Professor
BSc, University of Leeds, 1979-1983
PhD, University of London - Birkbeck College, 1983-1986
Postdoc, Massachusetts Institute of Technology, 1986-1989
Postdoc, Commissariat à L’Energie Atomique, 1989-1990
Researche de Recherche, CNRS, Institut Pasteur, 1990-1991
| Address | The Hospital for Sick Children Peter Gilgan Centre for Research and Learning (PGCRL) Room 20-9715 686 Bay Street Toronto, ON M5G 0A4 |
| Lab | Howell Lab |
| Lab Phone | 416-813-5354 or 416-813-5930 |
| Office Phone | 416-813-5378 |
| howell@sickkids.ca |
Research Interests
Our research activities are focused on understanding at the molecular and cellular level biological processes involved in microbial pathogenesis. The insights we gain from our fundamental research are used to develop novel strategies and treatments for bacterial and fungal biofilm related infections.
Innovations
Dr. Howell’s research has led to the identification of several glycosyl hydrolases that are capable of both preventing biofilm formation and degrading preformed biofilms. These enzymes are being developed as combination therapies for the treatment of a range of bacterial and fungal infections, and as disinfectants and coatings for cleaning and protecting surfaces.
Social Media
In the News
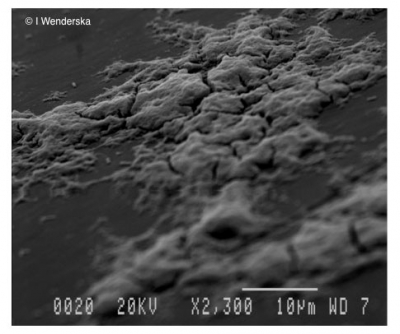
Research Lab

Wrinkly colony morphology of Pseudomonas aeruginosa Pel polysaccharide over expressing strain
Lab Members
Senior Scientist: Dr. P. Lynne Howell
Lab Manager: Dr. Stephanie Tammam
Lab Coordinator: Mr. Patrick Yip
Tech II: Ms. Ira Lacdao
Research Associates:
Dr. Roland Pfoh
Post-doctoral Fellows:
Dr. Perrin Baker
Dr. Kristin Low
Dr. Shane Caldwell
Graduate Students:
Ms. Natalie Bamford
Ms. Andreea Gheorghita
Ms. Lindsey Marmont
Mr. Matthew McCallum
Ms. Erum Razvi
Mr. Greg Whitfield
Learn more: Howell Lab
Research Description
Microbial Biofilm Formation: From Molecular Mechanisms to Therapeutics
Molecular Mechanisms
A microbial biofilm is a surface attached community of microbial cells encased in a self-produced polymeric matrix. Biofilms grow on any moist biotic or abiotic surface, and provide a number of environmental advantages such as protecting microbes from the host immune response, and conferring tolerance to antibiotics and detergents. Established biofilms are extremely difficult to eradiate and cost the health care systems in the US and Canada upwards of $25 billion annually to treat. They are responsible for greater than 65-80% of all chronic bacterial infections.
Using a multidisciplinary approach, which combines in vivo microbiological studies with structural biology, biophysical and biochemical techniques, we want to understand at the molecular level how two critical fundamental processes occur.
- How are exopolysaccharides – a major component of the biofilm matrix – synthesized, chemically modified, and exported from the cell; and
- What is required for the assembly and function of type IV pili – key virulence factors used by Pseudomonas aeruginosa to establish infection and for biofilm formation.
Therapeutics
Our research on the biosynthetic machinery of various exopolysaccharides has led to the identification of several glycoside hydrolases that are capable of both preventing biofilm formation and degrading preformed biofilms. These enzymes are effective at low nano-molar concentration and are being developed as combination therapies for the treatment of a range of bacterial and fungal infections, and as disinfectants and coatings for cleaning and protecting surfaces.
In addition, as modification of exopolysaccharides by the removal or addition of acetyl groups is crucial for the ability of many pathogenic organisms to initiate and maintain infection, we are employing high-throughput screening techniques and structure based drug design methods to identify small molecules that modulate the de-N-acetylation state of several different polymers involved in biofilm formation.
Exopolysaccharide polymerization, modification and export.
Exopolysaccharides are a major component of many microbial biofilms. These long sugar polymers are often found both associated with the bacterial or fungal cell surface and in a secreted form. Exopolysaccharide biosynthesis is a multistep process that requires the polymerization of nucleotide-sugar precursors, and transport across either the cytoplasmic or plasma membrane. In the case of gram-negative bacteria the polymer also needs to transverse the periplasm and be exported from the cell. The properties of the polymer are frequently modulated by chemically modification, which can occur either in the periplasm or extracellular space.
Ongoing projects seek to determine the molecular mechanisms involved in the biosynthesis of:
• alginate;
• Pel polysaccharide;
• Psl polysaccharide;
• poly β-1,6-N-acetyl-D-glucosamine (PNAG); and
• galactosaminogalactan (GAG).
Assembly of Pseudomonas aeruginosa Type IV pili
Type IV pili (T4P) are strong flexible filaments that mediate attachment to both living and artificial surfaces. They are also involved in bacteriophage adsorption, DNA uptake, biofilm initiation and development, and “twitching motility”, a unique form of surface-associated movement whereby the bacteria pull themselves rapidly towards or along a surface by retracting their T4P. Bacteria lacking T4P cannot adhere to surfaces and are therefore avirulent.
Regulation and assembly of the T4P is a complex process involving over 50 proteins. We are interested in understanding how a subset of the proteins in this molecular machine interact with each other to assembly, extend and retract the pilus.
Awards & Distinctions
2013-2020 — Canada Research Chair in Structural Biology Research Award Canadian Government
2006-2013 — Canada Research Chair in Structural Biology Research Award Canadian Government
2001-2006 — Investigator Research Award Canadian Institutes of Health Research
1983-1986 — Studentship Research Award Science and Engineering Research Council of Great Britain
Courses Taught
JBB2025H Protein Crystallography
BCH473Y Advanced Research Project in Biochemistry
Publications
View all publications on PubMed
Structure of the Pseudomonas aeruginosa Type IVa Pilus Secretin at 7.4 Å.
Koo J, Lamers RP, Rubinstein JL, Burrows LL, Howell PL.
Structure. 2016 Oct 4;24(10):1778-1787. doi: 10.1016/j.str.2016.08.007. Read
Exopolysaccharide biosynthetic glycoside hydrolases can be utilized to disrupt and prevent Pseudomonas aeruginosa biofilms.
Perrin Baker, Preston J. Hill, Brendan D. Snarr, Noor Alnabelseya, Matthew J. Pestrak, Mark J. Lee, Laura K. Jennings, John Tam, Roman A. Melnyk, Matthew R. Parsek, Donald C. Sheppard, Daniel J. Wozniak andP. Lynne Howell.
Science Advances 20 May 2016: Vol. 2, no. 5, e1501632. DOI: 10.1126/sciadv.1501632 Read
Modification and periplasmic translocation of the biofilm exopolysaccharide Poly-β-1,6-N-acetyl-D-glucosamine.
J. Little, G. Li, C. Ing, B.R. DiFrancesco, N. Bamford, H. Robinson, M. Nitz, R. Pomès, P. L. Howell.
Proc. Nat. Acad. Sci., July 3, 2014. Read
P. aeruginosa SGNH hydrolase-like proteins AlgJ and AlgX have similar topology but separate and distinct roles in alginate acetylation.
P. aeruginosa SCNH hydrolase-like proteins AlgJ and AlgX have similar topology but separate and distinct roles in alginate acetylation.
Baker P, Ricer T, Moynihan PJ, Kitova EN, Walvoort MT, Little DJ, Whitney JC, Dawson K, Weadge JT, Robinson H, Ohman DE, Codée JD, Klassen JS, Clarke AJ, Howell PL.
PLoS Pathog. 2014 10(8):e1004334. Read
Characterization of the PilN, PilO and PilP type IVa pilus subcomplex.
Tammam, P. Sundaram, J. Koo, L. Sampaleanu, M. Ayers, A. Chong, J. Forman-Kay, L.L. Burrows, P. L. Howell.
Mol. Micro. 82(6), 1496-1514, 2011. Read